library(tidyverse)
#> ── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
#> ✔ dplyr 1.2.0 ✔ readr 2.2.0
#> ✔ forcats 1.0.1 ✔ stringr 1.6.0
#> ✔ ggplot2 4.0.2 ✔ tibble 3.3.1
#> ✔ lubridate 1.9.5 ✔ tidyr 1.3.2
#> ✔ purrr 1.2.1
#> ── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
#> ✖ dplyr::filter() masks stats::filter()
#> ✖ dplyr::lag() masks stats::lag()
#> ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errorsIntroduction to the tidyverse
Scientific workflows: Tools and Tips 🛠️
2024-02-15
What is this lecture series?
Scientific workflows: Tools and Tips 🛠️
📅 Every 3rd Thursday 🕓 4-5 p.m. 📍 Webex
- One topic from the world of scientific workflows
- Material provided online
- If you don’t want to miss a lecture
What is the tidyverse?

The tidyverse is an opinionated collection of R packages designed for data science. All packages share an underlying design philosophy, grammar, and data structures.
www.tidyverse.org
Tidyverse core packages
Why the tidyverse?

Basic idea: Make data analysis efficient and intuitive
- Write code that is easy to read, write and maintain
- More time for data interpretation
- Tidyverse is actively developed and has a large community.
Today

Overview of
- most important packages and functions
- how they work together in a data analysis workflow
- underlying principles of the tidyverse
I can’t show you everything. But the tidyverse has an excellent documentation and cheatsheets.
Which tidyverse packages do you use?
Head over to the menti poll

Data analysis with the tidyverse




Getting the tidyverse
Install all tidyverse packages with
Load all core tidyverse packages with
You can also load packages individually - But don’t do both!
Tibbles from tibble
The underlying data structure

What are tibbles?

Tibbles are
a modern reimagining of the data frame. Tibbles are designed to be (as much as possible) drop-in replacements for data frames.
(Wickham, Advanced R)
Tibbles have the same basic properties as data frames
Everything that you can do with data frames, you can do with tibbles
Main advantage: Tibbles print much nicer to the console
Have a look at this book chapter for a full list of the differences between data frames and tibbles and the advantages of using tibbles.
How to create tibbles?

- Tidyverse functions return tibbles by default
- Create with
tibblefunction (equivalent todata.frame)
How to create tibbles?

Creating a tibble
Creating a data frame
data.frame(
x = 1:26,
y = letters,
z = factor(LETTERS)
)
#> x y z
#> 1 1 a A
#> 2 2 b B
#> 3 3 c C
#> 4 4 d D
#> 5 5 e E
#> 6 6 f F
#> 7 7 g G
#> 8 8 h H
#> 9 9 i I
#> 10 10 j J
#> 11 11 k K
#> 12 12 l L
#> 13 13 m M
#> 14 14 n N
#> 15 15 o O
#> 16 16 p P
#> 17 17 q Q
#> 18 18 r R
#> 19 19 s S
#> 20 20 t T
#> 21 21 u U
#> 22 22 v V
#> 23 23 w W
#> 24 24 x X
#> 25 25 y Y
#> 26 26 z ZImport data with readr
Read in your files
Most important readr functions

read_csvto read comma delimited filesread_tsvto read tab delimited filesread_delimto read files with any delimiter
All read_* functions return a tibble.
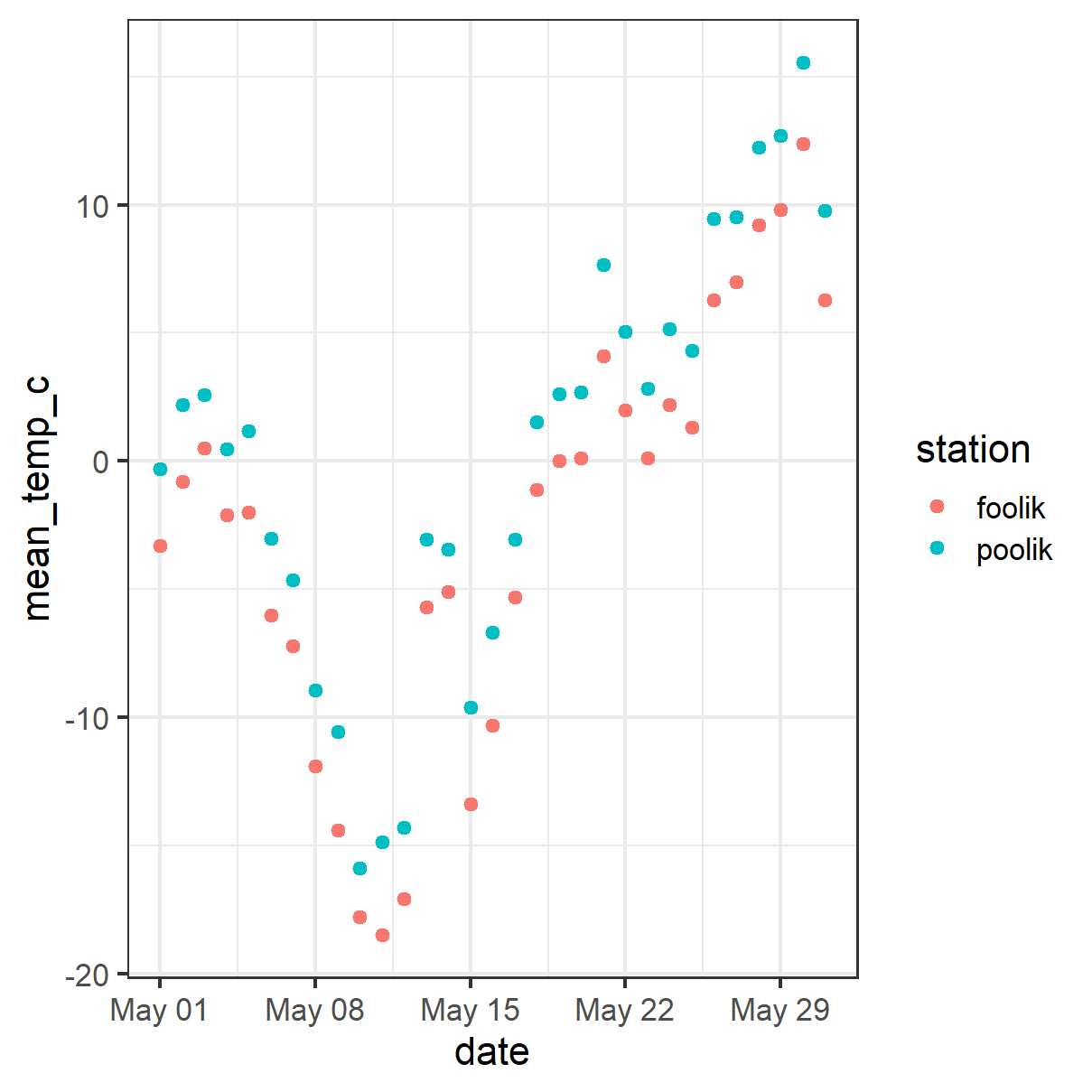
Example with arctic temperature data

Data modified from lterdatasampler
arc_weather <- read_csv(file = "data/arc_weather.csv")
arc_weather
#> # A tibble: 365 × 3
#> date foolik_mean_temp_c poolik_mean_temp_c
#> <date> <dbl> <dbl>
#> 1 1988-06-01 8.4 11.8
#> 2 1988-06-02 6 8.90
#> 3 1988-06-03 5.8 8.14
#> 4 1988-06-04 1.8 3.95
#> 5 1988-06-05 6.8 9.81
#> 6 1988-06-06 5.2 8.74
#> 7 1988-06-07 2.2 6.11
#> 8 1988-06-08 9.4 12.1
#> 9 1988-06-09 13.1 16.5
#> 10 1988-06-10 17.7 20.9
#> # ℹ 355 more rowsAdvantages of readr

Faster than
read.table/read.csvBetter defaults than base R (e.g. guessing of data types)
Returns tibbles
Tidyverse packages for other data types:
readxlfor Excel fileshavenfor SPSS, SAS, and Stata filesgooglesheets4for Google sheets- …
Tidy data with tidyr
Reorganize the data

What is tidy data?
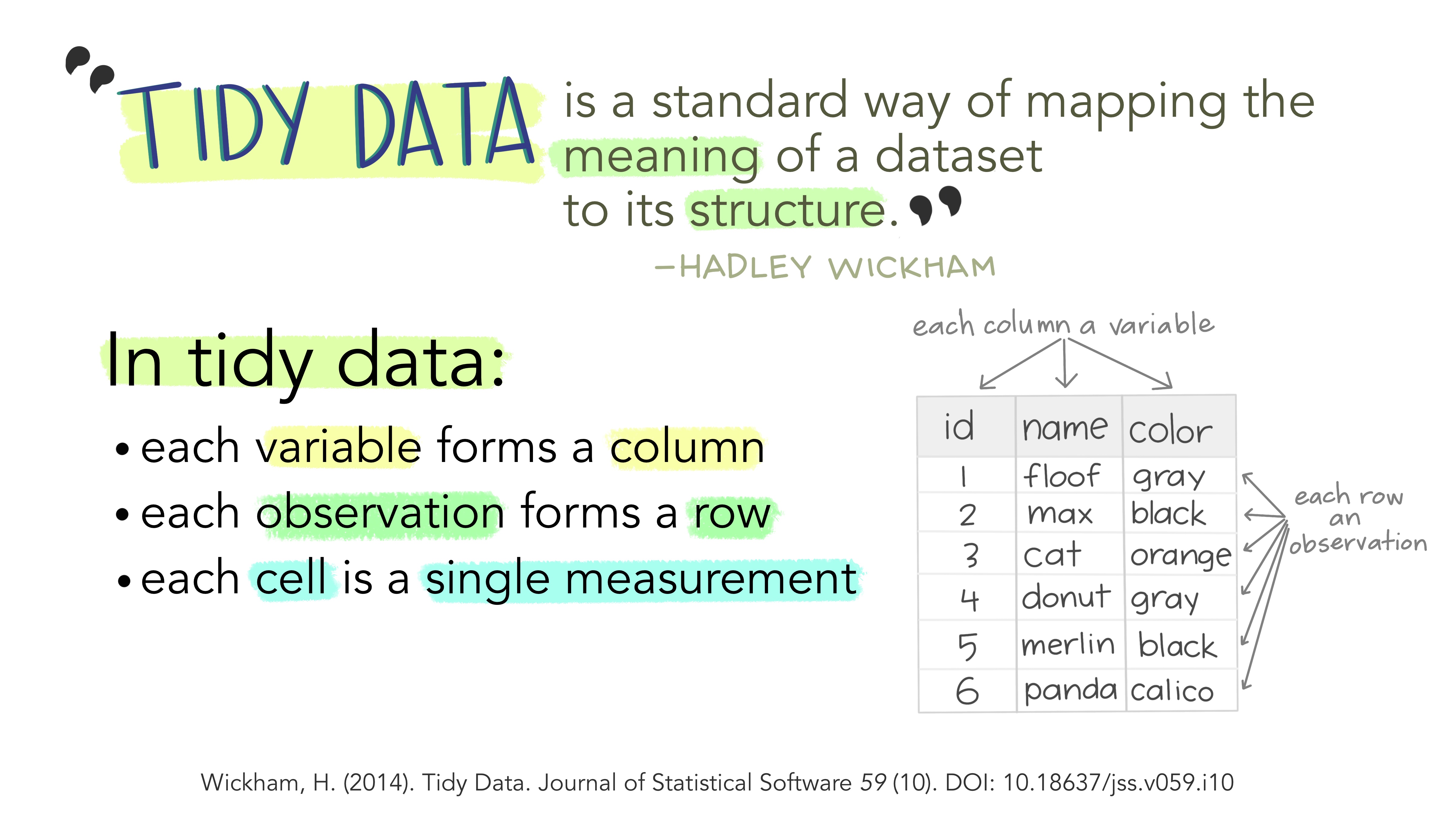

What is non-tidy data?
Tidy
| id | name | color |
|---|---|---|
| 1 | floof | gray |
| 2 | max | black |
| 3 | cat | orange |
| 4 | donut | gray |
| 5 | merlin | black |
| 6 | panda | calico |
Non-tidy
| floof | max | cat | donut | merlin | panda |
|---|---|---|---|---|---|
| gray | black | orange | gray | black | calico |
| gray | black | orange | calico |
|---|---|---|---|
| floof | max | cat | panda |
| donut | merlin |
Sometimes raw data is non-tidy because its structure is optimized for data entry or viewing rather than analysis.
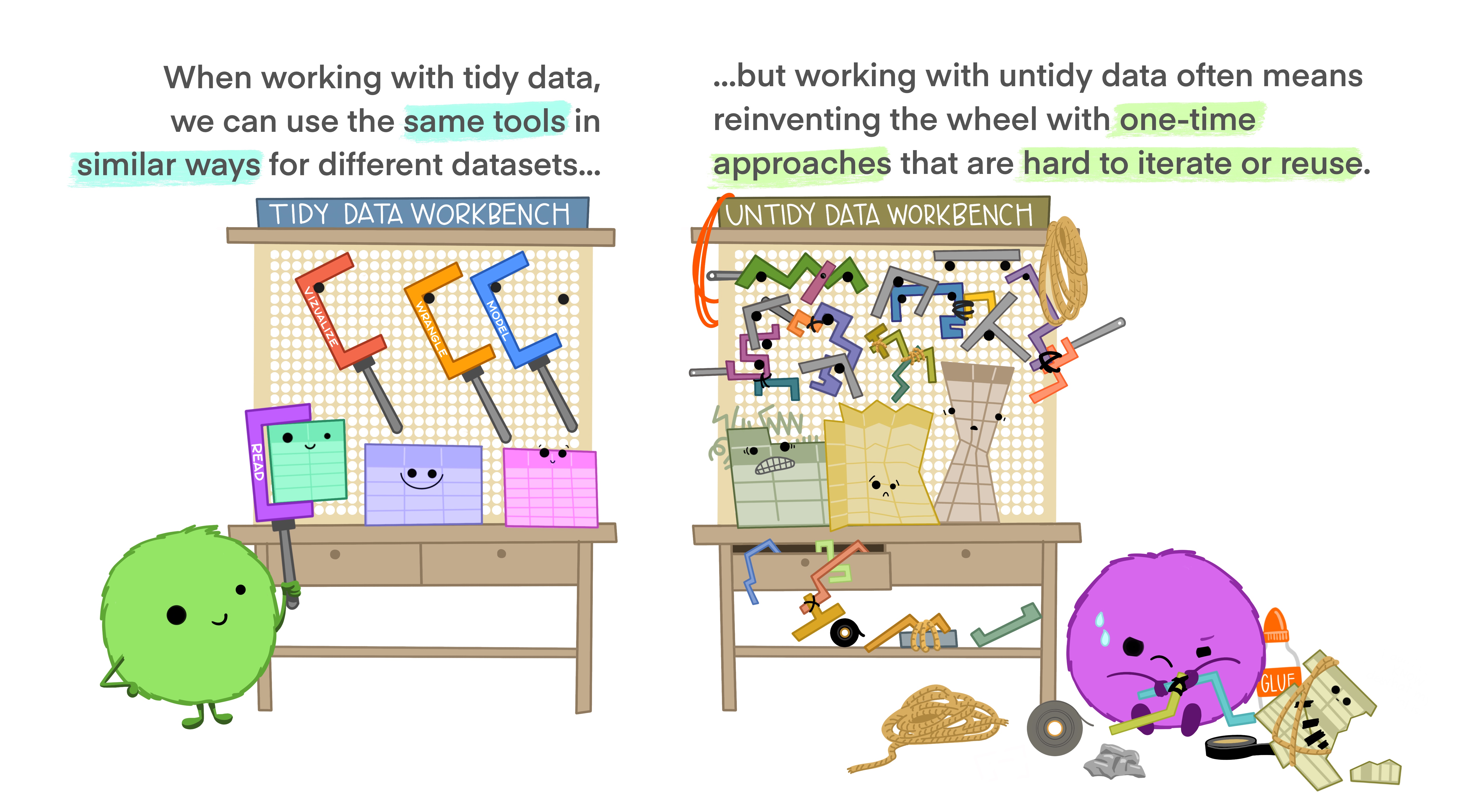
Advantages of tidy data
The main advantages of tidy data is that the tidyverse packages are built to work with it.
Tidy data with tidyr

Tidyrreorganizes the data but does not change values- Most important functionality:
- Pivoting:
pivot_longerandpivot_wider - Splitting and combining columns:
separate_wider_delim/position/regexandunite - Handle missing values:
drop_na,replace_na,complete
- Pivoting:
Example

What is not tidy about our weather data?
arc_weather
#> # A tibble: 365 × 3
#> date foolik_mean_temp_c poolik_mean_temp_c
#> <date> <dbl> <dbl>
#> 1 1988-06-01 8.4 11.8
#> 2 1988-06-02 6 8.90
#> 3 1988-06-03 5.8 8.14
#> 4 1988-06-04 1.8 3.95
#> 5 1988-06-05 6.8 9.81
#> 6 1988-06-06 5.2 8.74
#> 7 1988-06-07 2.2 6.11
#> 8 1988-06-08 9.4 12.1
#> 9 1988-06-09 13.1 16.5
#> 10 1988-06-10 17.7 20.9
#> # ℹ 355 more rows- Each row has multiple observations
- Variables are split across multiple columns
Example

This can be solved with pivot_longer
pivot_longer(arc_weather,
cols = c(foolik_mean_temp_c, poolik_mean_temp_c))
#> # A tibble: 730 × 3
#> date name value
#> <date> <chr> <dbl>
#> 1 1988-06-01 foolik_mean_temp_c 8.4
#> 2 1988-06-01 poolik_mean_temp_c 11.8
#> 3 1988-06-02 foolik_mean_temp_c 6
#> 4 1988-06-02 poolik_mean_temp_c 8.90
#> 5 1988-06-03 foolik_mean_temp_c 5.8
#> 6 1988-06-03 poolik_mean_temp_c 8.14
#> 7 1988-06-04 foolik_mean_temp_c 1.8
#> 8 1988-06-04 poolik_mean_temp_c 3.95
#> 9 1988-06-05 foolik_mean_temp_c 6.8
#> 10 1988-06-05 poolik_mean_temp_c 9.81
#> # ℹ 720 more rowsExample

You can also directly name the new columns:
arc_weather <- pivot_longer(arc_weather,
cols = c(foolik_mean_temp_c, poolik_mean_temp_c),
names_to = "station",
values_to = "mean_temp_c")
arc_weather
#> # A tibble: 730 × 3
#> date station mean_temp_c
#> <date> <chr> <dbl>
#> 1 1988-06-01 foolik_mean_temp_c 8.4
#> 2 1988-06-01 poolik_mean_temp_c 11.8
#> 3 1988-06-02 foolik_mean_temp_c 6
#> 4 1988-06-02 poolik_mean_temp_c 8.90
#> 5 1988-06-03 foolik_mean_temp_c 5.8
#> 6 1988-06-03 poolik_mean_temp_c 8.14
#> 7 1988-06-04 foolik_mean_temp_c 1.8
#> 8 1988-06-04 poolik_mean_temp_c 3.95
#> 9 1988-06-05 foolik_mean_temp_c 6.8
#> 10 1988-06-05 poolik_mean_temp_c 9.81
#> # ℹ 720 more rowsTransform data with dplyr
How to actually change values
Transform data with dplyr

Data cleaning, adding new columns, summarizing, …
Depending on the data type in combination with
stringrfor character columnslubridatefor date-time columnsforcatsfor factor columns
Data cleaning and filtering

select picks variables (columns) based on their names
select(arc_weather,
date, mean_temp_c)
#> # A tibble: 730 × 2
#> date mean_temp_c
#> <date> <dbl>
#> 1 1988-06-01 8.4
#> 2 1988-06-01 11.8
#> 3 1988-06-02 6
#> 4 1988-06-02 8.90
#> 5 1988-06-03 5.8
#> 6 1988-06-03 8.14
#> 7 1988-06-04 1.8
#> 8 1988-06-04 3.95
#> 9 1988-06-05 6.8
#> 10 1988-06-05 9.81
#> # ℹ 720 more rowsfilter picks observations (rows) based on their values
filter(arc_weather,
station == "foolik_mean_temp_c")
#> # A tibble: 365 × 3
#> date station mean_temp_c
#> <date> <chr> <dbl>
#> 1 1988-06-01 foolik_mean_temp_c 8.4
#> 2 1988-06-02 foolik_mean_temp_c 6
#> 3 1988-06-03 foolik_mean_temp_c 5.8
#> 4 1988-06-04 foolik_mean_temp_c 1.8
#> 5 1988-06-05 foolik_mean_temp_c 6.8
#> 6 1988-06-06 foolik_mean_temp_c 5.2
#> 7 1988-06-07 foolik_mean_temp_c 2.2
#> 8 1988-06-08 foolik_mean_temp_c 9.4
#> 9 1988-06-09 foolik_mean_temp_c 13.1
#> 10 1988-06-10 foolik_mean_temp_c 17.7
#> # ℹ 355 more rowsCombine filter with other packages

lubridate to filter by month
# Filter rows from May
filter(arc_weather,
month(date) == 5)
#> # A tibble: 62 × 3
#> date station mean_temp_c
#> <date> <chr> <dbl>
#> 1 1989-05-01 foolik_mean_temp_c -3.3
#> 2 1989-05-01 poolik_mean_temp_c -0.312
#> 3 1989-05-02 foolik_mean_temp_c -0.8
#> 4 1989-05-02 poolik_mean_temp_c 2.20
#> 5 1989-05-03 foolik_mean_temp_c 0.5
#> 6 1989-05-03 poolik_mean_temp_c 2.59
#> 7 1989-05-04 foolik_mean_temp_c -2.1
#> 8 1989-05-04 poolik_mean_temp_c 0.457
#> 9 1989-05-05 foolik_mean_temp_c -2
#> 10 1989-05-05 poolik_mean_temp_c 1.19
#> # ℹ 52 more rowsstringr to filter characters
# Filter rows where the station contains "foolik"
filter(arc_weather,
str_detect(station, "foolik"))
#> # A tibble: 365 × 3
#> date station mean_temp_c
#> <date> <chr> <dbl>
#> 1 1988-06-01 foolik_mean_temp_c 8.4
#> 2 1988-06-02 foolik_mean_temp_c 6
#> 3 1988-06-03 foolik_mean_temp_c 5.8
#> 4 1988-06-04 foolik_mean_temp_c 1.8
#> 5 1988-06-05 foolik_mean_temp_c 6.8
#> 6 1988-06-06 foolik_mean_temp_c 5.2
#> 7 1988-06-07 foolik_mean_temp_c 2.2
#> 8 1988-06-08 foolik_mean_temp_c 9.4
#> 9 1988-06-09 foolik_mean_temp_c 13.1
#> 10 1988-06-10 foolik_mean_temp_c 17.7
#> # ℹ 355 more rowsChange and add values

Add columns with mutate
# Add column with temperature in K
mutate(arc_weather,
mean_temp_k = mean_temp_c + 274.15)
#> # A tibble: 730 × 4
#> date station mean_temp_c mean_temp_k
#> <date> <chr> <dbl> <dbl>
#> 1 1988-06-01 foolik_mean_temp_c 8.4 283.
#> 2 1988-06-01 poolik_mean_temp_c 11.8 286.
#> 3 1988-06-02 foolik_mean_temp_c 6 280.
#> 4 1988-06-02 poolik_mean_temp_c 8.90 283.
#> 5 1988-06-03 foolik_mean_temp_c 5.8 280.
#> 6 1988-06-03 poolik_mean_temp_c 8.14 282.
#> 7 1988-06-04 foolik_mean_temp_c 1.8 276.
#> 8 1988-06-04 poolik_mean_temp_c 3.95 278.
#> 9 1988-06-05 foolik_mean_temp_c 6.8 281.
#> 10 1988-06-05 poolik_mean_temp_c 9.81 284.
#> # ℹ 720 more rowsChange and add values

Change the values of existing columns with mutate
# Remove _mean_temp_c from station
arc_weather <- mutate(arc_weather,
station = str_remove(station, "_mean_temp_c"))
arc_weather
#> # A tibble: 730 × 3
#> date station mean_temp_c
#> <date> <chr> <dbl>
#> 1 1988-06-01 foolik 8.4
#> 2 1988-06-01 poolik 11.8
#> 3 1988-06-02 foolik 6
#> 4 1988-06-02 poolik 8.90
#> 5 1988-06-03 foolik 5.8
#> 6 1988-06-03 poolik 8.14
#> 7 1988-06-04 foolik 1.8
#> 8 1988-06-04 poolik 3.95
#> 9 1988-06-05 foolik 6.8
#> 10 1988-06-05 poolik 9.81
#> # ℹ 720 more rowsSummarize data

summarize will collapse the data to a single row
Summarizing data by group

summarize in combination with group_by will collapse the data to a single row per group
group_bycan be used for any dplyr function to perform the operation by group
The pipe |>
Data transformation often requires multiple operations in sequence.
The pipe operator |> helps to keep these operations clear and readable.
- You may also see
%>%from themagrittrpackage (tidyverse) - Turn on the native R pipe
|>in Tools -> Global Options -> Code
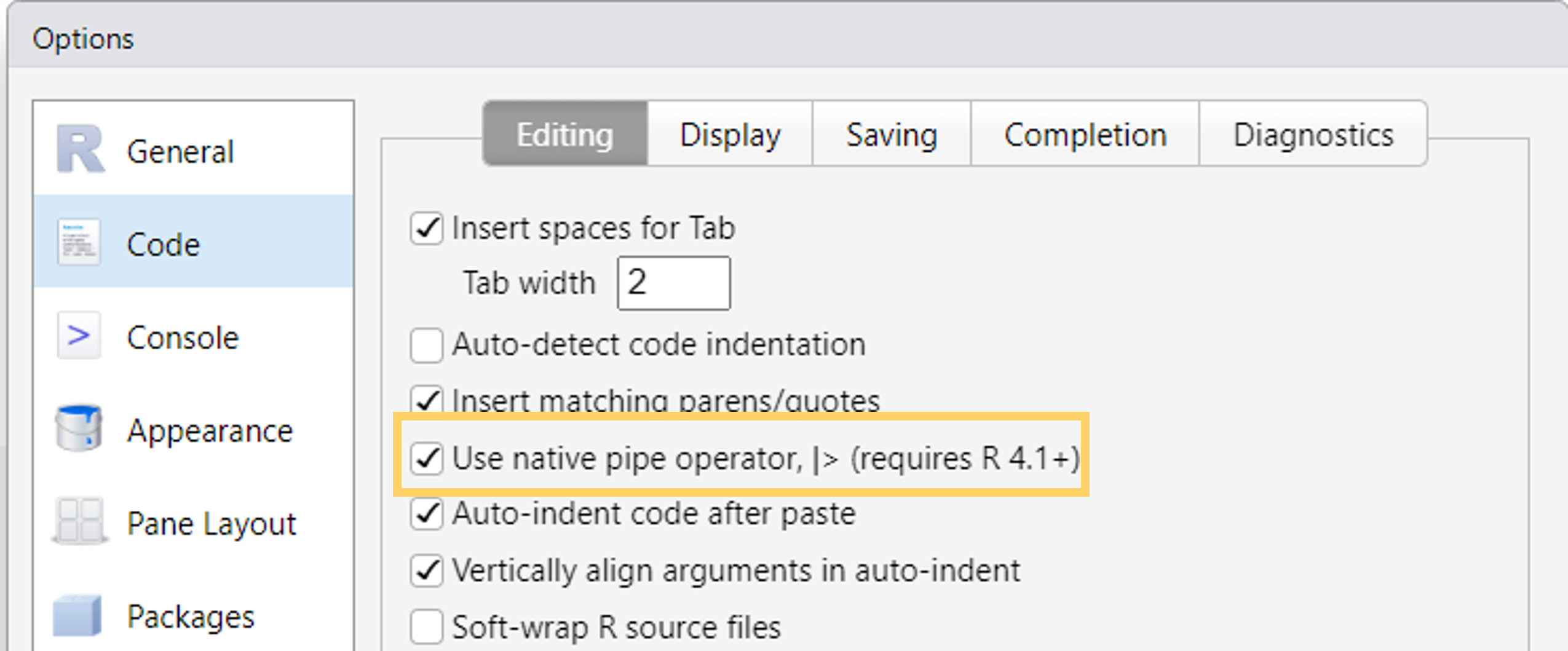
The pipe |>
Let’s look at an example without the pipe:
The pipe operator makes it very easy to combine these operations
You can read from top to bottom and interpret the |> as an “and then do”.
The pipe |>
But what is happening?
The pipe is “pushing” the result of one line into the first argument of the function from the next line.
Piping works perfectly with the tidyverse functions because they are designed to return a tibble and take a tibble as first argument.
Tip
Use the keyboard shortcut Ctrl/Cmd + Shift + M to insert |>
Visualize data with ggplot2
Quick exploratory plots and data masterpieces
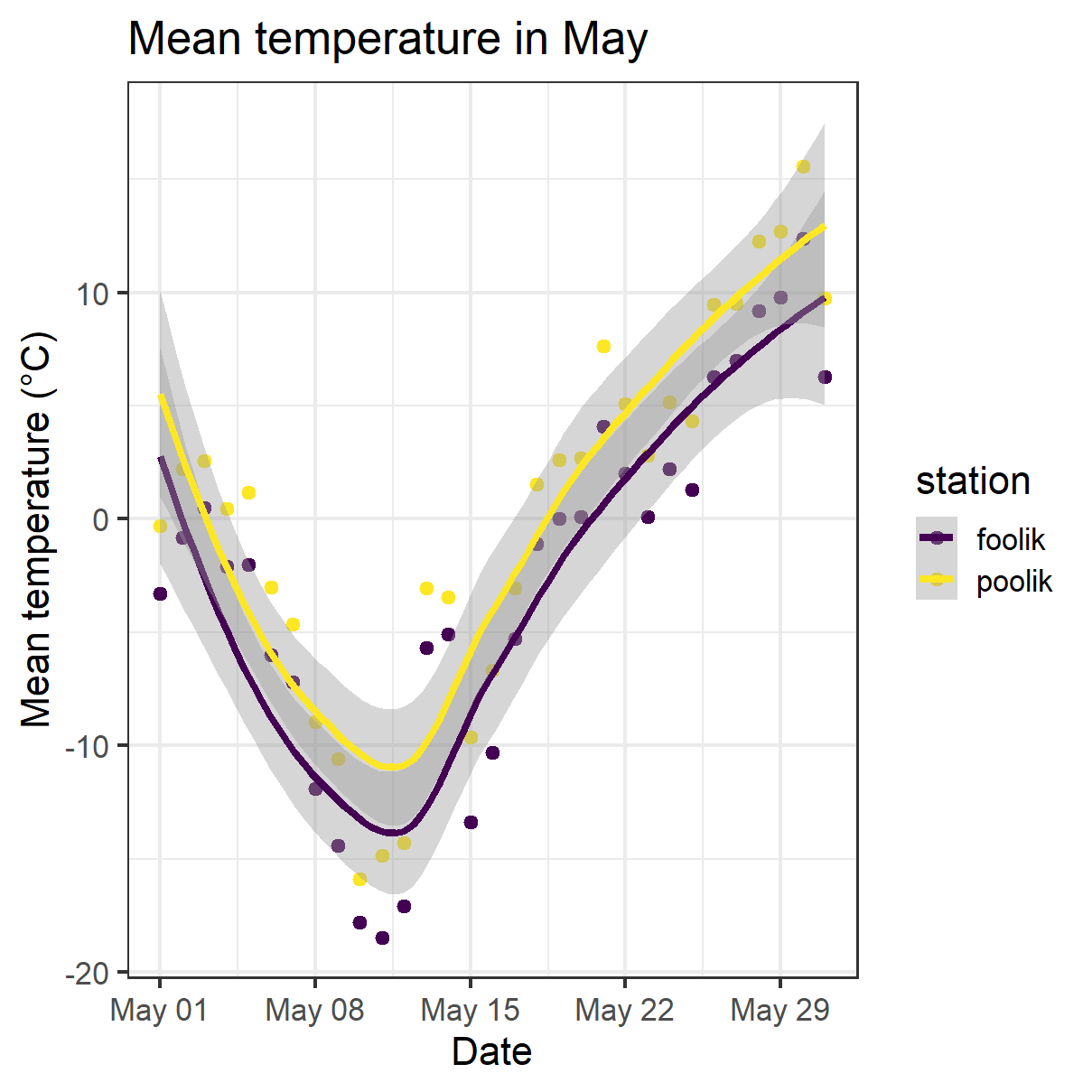
A ggplot showcase

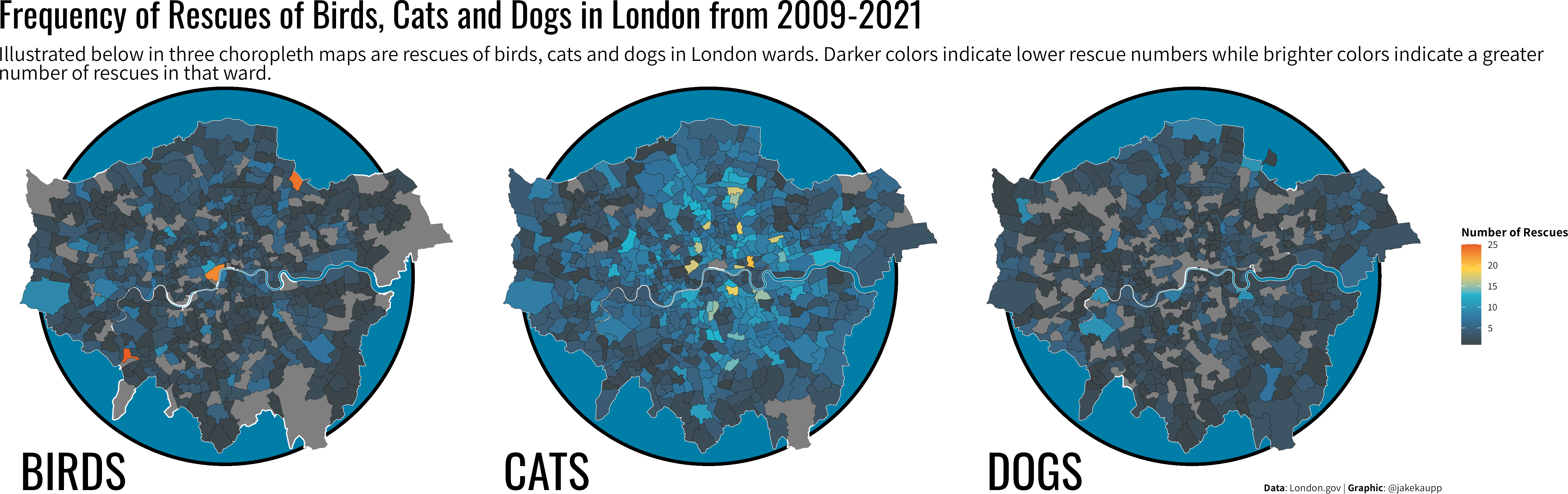
A ggplot showcase

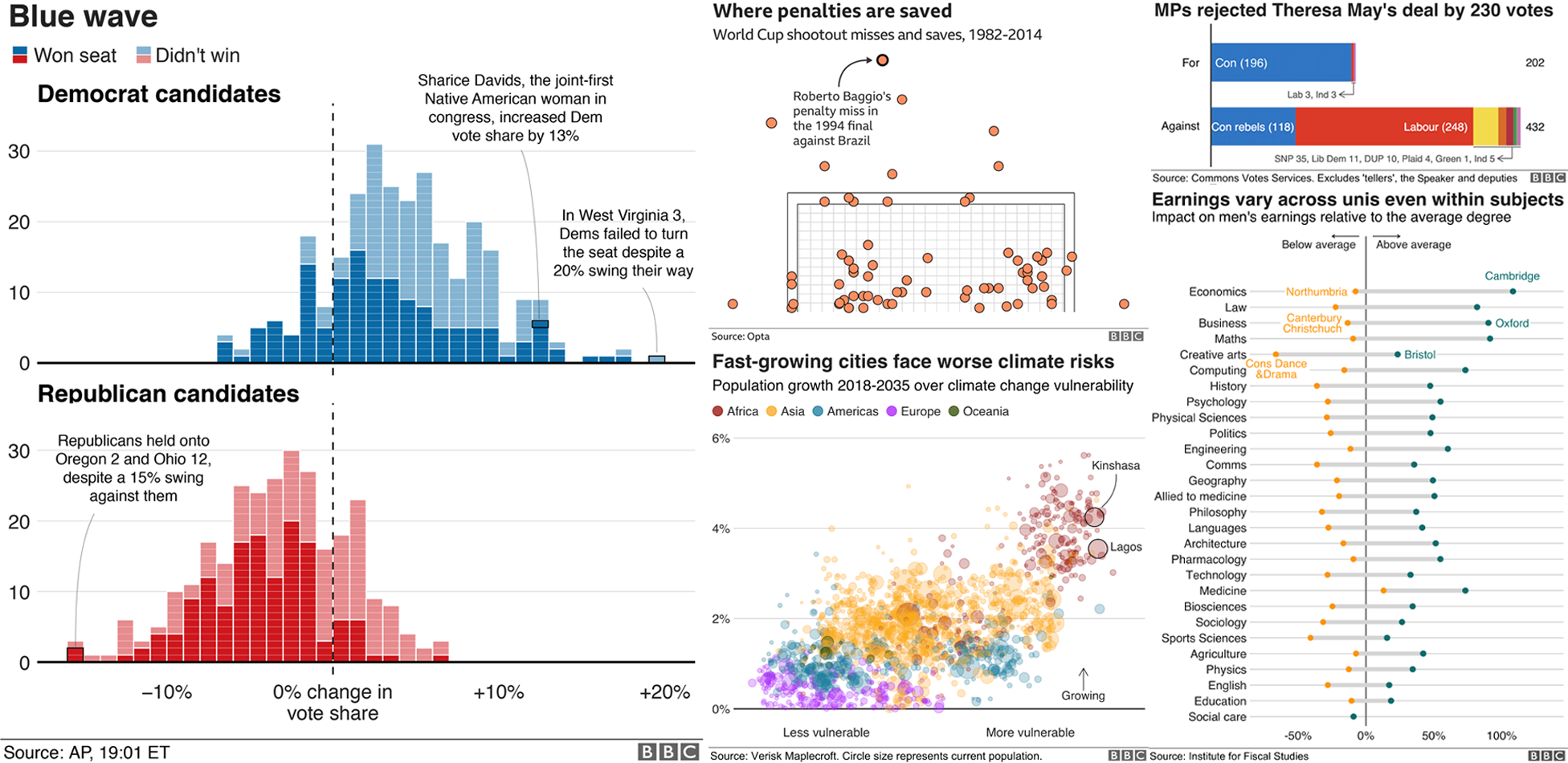
A ggplot showcase

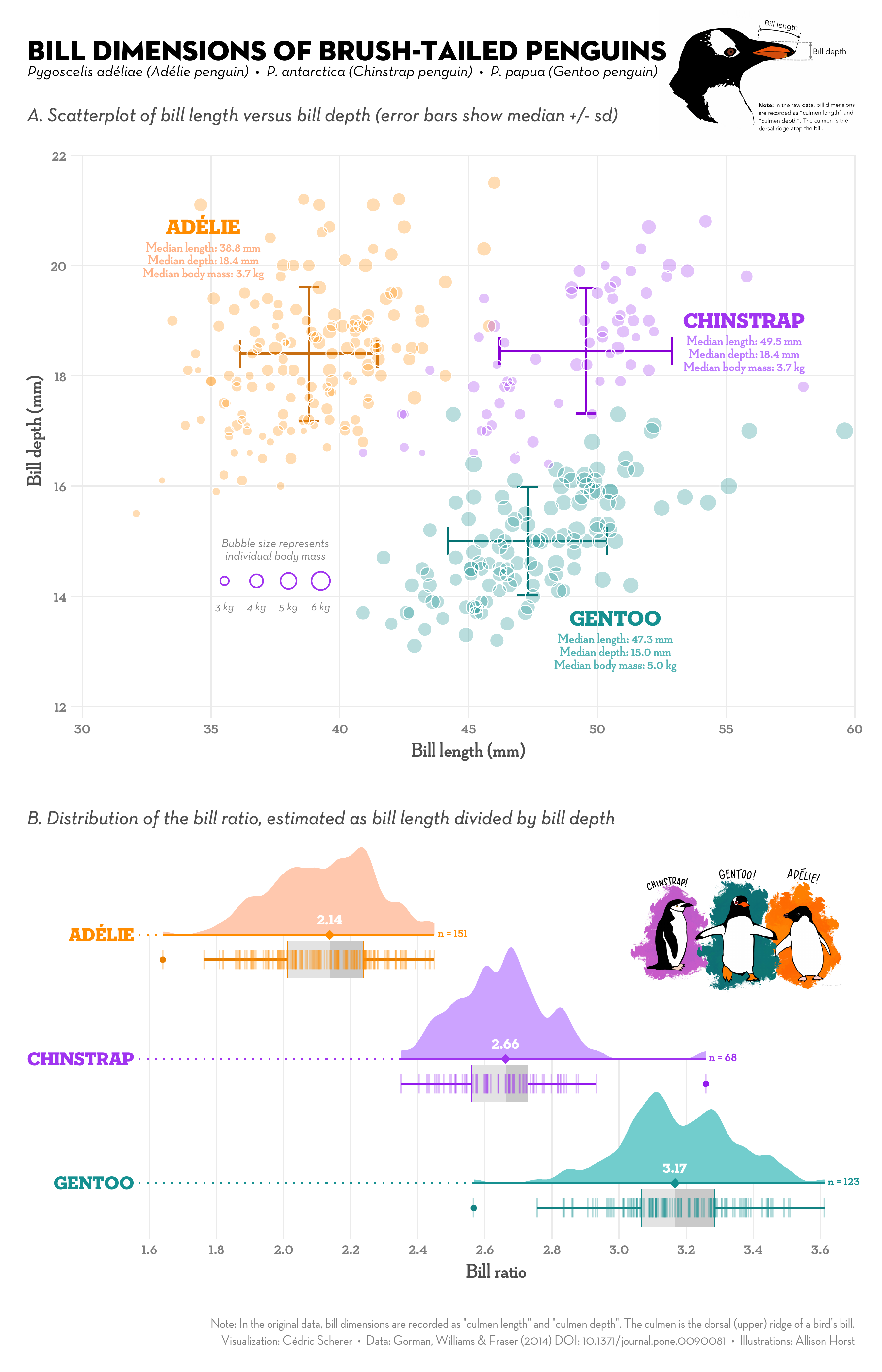
Advantages of ggplot

- Consistent grammar
- Flexible structure allows you to produce any type of plots
- Highly customizable appearance (themes)
- Many extension packages that provide additional plot types, themes, colors, animation, …
- Find a list of ggplot extensions here
- Perfect package for exploratory data analysis and beautiful plots
Basic ggplot

Basic idea: Stack layers of the plot on top of each other with +
Basic ggplot

- Gplot offers many more customization options and layers
- Works well with the pipe
|>
Functional programming with purrr

Functional programming with purrr

- Apply functions to multiple elements of a list/vector/…
- Replace e.g. for loops/apply-functions
- Often
purrrfunctions are more intuitive - See here for a full comparison between base R functions and purr functions
- Most important function:
map- Comes in different versions, depending on the input and the desired output
A simple example

Draw 4 numbers from normal distribution with means 1, 2, 3, 4.
Do it by hand
# Do it by hand
set.seed(123)
rnorm(n = 4, mean = 1, sd = 1)
#> [1] 0.4395244 0.7698225 2.5587083 1.0705084
rnorm(n = 4, mean = 2, sd = 1)
#> [1] 2.1292877 3.7150650 2.4609162 0.7349388
rnorm(n = 4, mean = 3, sd = 1)
#> [1] 2.313147 2.554338 4.224082 3.359814
rnorm(n = 4, mean = 4, sd = 1)
#> [1] 4.400771 4.110683 3.444159 5.786913Different versions of the map function

These are just some examples, there are many more options:
| Output | Input | Purr function |
|---|---|---|
| List | 1 Vector | map |
| List | 2 Vectors | map2 |
| Vector of desired type | 1 Vector | map_lgl |
| Vector of desired type | 2 Vectors | map2_lgl |
| Data frame | 1 Vector | map_df |
| Data frame | 2 Vectors | map2_df |
A more complex example

Read in and analyse multiple files with the same structure at once.
#> [1] "foolik.csv" "poolik.csv"# Step 2: Read all files into a list of tibbles
arc_weather_files <- map(paths, read_csv)
arc_weather_files
#> [[1]]
#> # A tibble: 365 × 3
#> date station mean_temp_c
#> <date> <chr> <dbl>
#> 1 1988-06-01 foolik 8.4
#> 2 1988-06-02 foolik 6
#> 3 1988-06-03 foolik 5.8
#> 4 1988-06-04 foolik 1.8
#> 5 1988-06-05 foolik 6.8
#> 6 1988-06-06 foolik 5.2
#> 7 1988-06-07 foolik 2.2
#> 8 1988-06-08 foolik 9.4
#> 9 1988-06-09 foolik 13.1
#> 10 1988-06-10 foolik 17.7
#> # ℹ 355 more rows
#>
#> [[2]]
#> # A tibble: 365 × 3
#> date station mean_temp_c
#> <date> <chr> <dbl>
#> 1 1988-06-01 poolik 11.5
#> 2 1988-06-02 poolik 8.00
#> 3 1988-06-03 poolik 8.50
#> 4 1988-06-04 poolik 4.31
#> 5 1988-06-05 poolik 9.56
#> 6 1988-06-06 poolik 8.10
#> 7 1988-06-07 poolik 5.56
#> 8 1988-06-08 poolik 12.7
#> 9 1988-06-09 poolik 16.0
#> 10 1988-06-10 poolik 21.5
#> # ℹ 355 more rowsA more complex example

Read in and analyse multiple files with the same structure at once.
# Step 3: Apply a linear model to each station file,
# get the model summary of each model and extract the coefficients
arc_weather_files |> # Take the data
map(\(x) lm(mean_temp_c ~ date, data = x)) |> # Apply the same model to every tibbel in the list
map(summary) |> # Get the model summary of both models
map_dbl("r.squared") # Extract the r-squared from the model summaries
#> [1] 0.2018279 0.2000210Summary
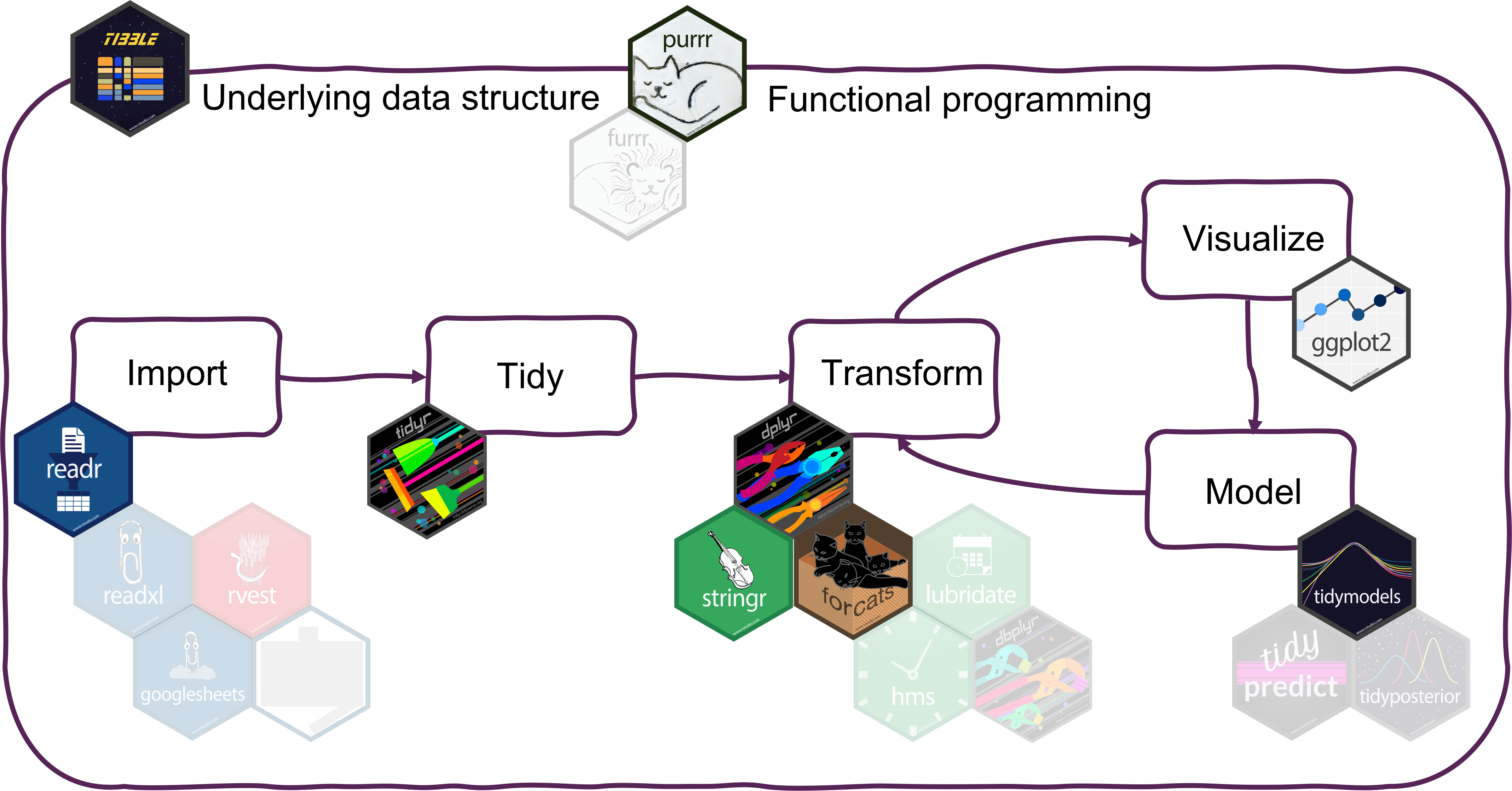
Summary
Tidyverse makes data analysis
- efficient
- intuitive
- consistent
Tidyverse code is more readable and easier to write & maintain.
This facilitates
📚 Learning
🤝 Collaboration
🔄 Reproducible research
Summary
Where to get started with the tidyverse?
- Check out the lecture website for links to documenation, cheatsheets and book

Next lecture
Semester break in March!
Topic t.b.a.
📅 18th April 🕓 4-5 p.m. 📍 Webex
🔔 Subscribe to the mailing list
📧 For topic suggestions and/or feedback send me an email
Thank you for your attention :)
Questions?
References
- Tidyverse website: find links to all package documentations
- Cheatsheets
- R for Data Science book: Learn data analysis with the tidyverse
Selina Baldauf // Tidyverse